Untangling RAS
An “undruggable” cancer-causing protein, 30 years of research and the national initiative that aims to finally make the connections
November 21, 2013 | Leigh MacMillan

Photograph by Retrofile/Getty Images
Picture a switchboard operator of old. There she sits, in front of a dizzying array of toggles and lights, ready to push buttons and stretch cords to connect incoming calls to just the right recipient. She might even listen in on the conversations from time to time.
The protein Ras is like that operator.
It sits inside a cell, ready to detect incoming signals and connect them to the next protein in a signaling cascade that controls cell behavior. The “conversations” that Ras connects can determine how fast a cell grows, whether it changes into a different cell type and even whether it lives or dies.
When Ras makes mistakes—when it fails to disconnect a “call”—it can cause cancer. Mutations that activate Ras cause cells to divide without a signal and to ignore cues telling them to die.
In fact, up to one-third of all human cancers have a mutation in one of the RAS genes (there are three of them), making RAS family members the most common cancer-causing genes. Activating RAS mutations are found in up to 90 percent of pancreatic cancers, 50 percent of colon cancers and 30 percent of lung cancers—three of the four deadliest cancers in the United States.
It has been three decades since mutations in RAS genes were first linked to human cancers. Studies since then have shown that turning off activated Ras reverses cancerous characteristics in cells and shrinks tumors in animal models—making Ras an attractive target for cancer therapy. Yet despite intensive investigation, there are no therapies directed at the faulty switch.
Perhaps the time is nigh.
A new national initiative—the Ras Project—aims to bring together the resources and expertise to finally crack the Ras riddle and discover drugs that block its cancer-causing ways.
“Increasing technology and increasing knowledge relating to the Ras pathway, as well as increasing awareness of its importance in the clinical setting, have all converged to make this the right time for this project,” says Frank McCormick, Ph.D., director of the Helen Diller Family Comprehensive Cancer Center at the University of California, San Francisco and associate dean of the UCSF School of Medicine.
McCormick, a longtime Ras researcher, is leading the national initiative, which is based at the Frederick National Laboratory for Cancer Research (SAIC-Frederick, Inc.), with oversight from the National Cancer Institute. The project launched this summer.
An old foe
The story of Ras goes back to the mid-1960s, when scientists Jennifer Harvey and Werner Kirsten isolated two viruses—Harvey murine sarcoma virus and Kirsten murine sarcoma virus—that caused tumors in rats and other rodents. The tumor-causing viral genes were named H-RAS and K-RAS, for their discoverers (H- and K-) and for their ability to cause rat sarcomas.
In the late 1970s, Edward Scolnick and his colleagues at the National Cancer Institute determined that the viral RAS genes were actually “normal” cellular genes that the viruses had picked up.
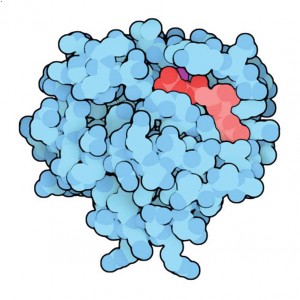
The name “Ras” is a shortened form of rat sarcomas, where the genes were originally found in the 1960s. When referring to the protein form, scientists use “Ras.” When referring to the gene, it’s “RAS.” Image from the RCSB PDB September 2008 Molecule of the Month feature by David Goodsell (doi:10.2210/rcsb_pdb/ mom_2008_9).
In 1982, the RAS genes were linked to human cancer (as opposed to virus-induced cancer in rodents) when three groups discovered that RAS genes possessing a single mutation could “transform” cultured cells and endow them with cancerous features. Genes with the ability to turn normal cells into cancer cells became known as “oncogenes;” the RAS genes were some of the first oncogenes identified.
The cancer-causing RAS mutations were subsequently found in tumor samples from patients and a third human RAS gene—N-RAS—was identified as being mutated in neuroblastoma.
Intense Ras-related research activity since 1982 established that Ras proteins function as biochemical switches and demonstrated how cellular signals flip Ras “on” to activate processes that control cell proliferation and death. Cancer-causing mutations leave Ras proteins stuck in the “on” position, prompting a search for strategies and drugs that turn Ras “off.”
Ras has been a tricky target.
No therapies turn Ras off, either directly or indirectly. And some of the newer targeted therapies—cetuximab for colorectal cancer, erlotinib for lung cancer, vemurafenib for melanoma—cannot be used in patients who have tumors with Ras mutations. These therapies may cause Ras-mutant cancers to progress more quickly.
“Of the various oncogene drivers that we know of, Ras is the one that has been resistant to attempts to target it,” McCormick says.
Teflon proteins
Proteins that make good drug targets usually have “pockets”—indentations—on the protein surface, where small chemicals can fit and bind tightly to alter protein activity.
Three-dimensional structures of the H-Ras protein, which has served as the model form of Ras, didn’t reveal any good pockets, says Stephen Fesik, Ph.D., Orrin H. Ingram II Professor of Cancer Research at Vanderbilt-Ingram Cancer Center.
This lack of suitable binding pockets puts Ras in a category of so-called “undruggable” targets.
“We call proteins like Ras Teflon proteins. Nothing sticks to them,” Fesik says.
Teflon proteins don’t faze Fesik.
Before he came to Vanderbilt to lead cancer drug discovery efforts, Fesik was vice president of Cancer Research at Abbott Laboratories, where he pioneered new methods to discover drugs for challenging targets.
“Our goal is to discover new drugs to treat cancer patients that will have a big impact,” Fesik says. “And we’re going after the proteins that pharmaceutical companies shy away from.”
One of those proteins is Ras.
Fesik’s team uses a method he developed called SAR by NMR (structure-activity relationships by nuclear magnetic resonance). The investigators use NMR methods to screen libraries of small chemical fragments for binding to a target protein. Then they use NMR or X-ray crystallography to determine how any chemical “hits” bind to the target.

Stephen Fesik, Ph.D. Photo by Daniel Dubois.
In the case of Ras, Fesik and his team decided to screen for drugs that bind to K-Ras, rather than to H-Ras.
Although study of H-Ras has dominated research efforts— and all three forms of Ras have high sequence similarity—K-Ras is the most frequently mutated form in human tumors (86 percent of all documented Ras mutations are in K-Ras; 11 percent are in N-Ras and 3 percent are in H-Ras).
Fesik and his colleagues first determined a structure of K-Ras using X-ray crystallography and confirmed that it, like H-Ras, had no apparent pockets. But they screened their chemical fragment library anyway.
“To our surprise we ended up finding molecules, very small molecules, that bound to K-Ras,” Fesik says. They found a small pocket that appeared, in both mutated and wild-type forms of K-Ras, in the presence of the chemical fragments.
The investigators further determined that their small molecules prevented formation of the activated state of Ras, keeping it in an inactive state. The studies were published in the journal Angewandte Chemie in May 2012.
In the meantime, the group has been optimizing the fragments to improve their binding characteristics as well as identifying fragments that bind to a second site on K-Ras and attempting to link fragments that bind at the two different sites.
The drug discovery advances positioned Fesik for a central role in the Ras Project—he is one of the leading investigators for the discovery of K-Ras inhibitors.
Hub-and-spokes
At the April annual meeting of the American Association for Cancer Research, Harold Varmus, M.D., director of the National Cancer Institute, announced his vision for a national Ras Project that would “finally after 30 years learn how to target the cancer cells … that are driven by mutations in RAS.”
The project, which UCSF’s McCormick is leading with SAIC-Frederick’s Chief Executive Officer David Heimbrook, Ph.D., got the go-ahead from two NCI advisory boards in June. About $10 million per year of SAIC-Frederick’s budget will be re-allocated to support the Ras Project, and McCormick will devote about half of his time to the effort.
McCormick describes the project as having a central “hub”— the Frederick National Laboratory—surrounded by “spokes” reaching out to academic and biopharmaceutical industry partners.
“This is such a wonderful opportunity for me personally, and it’s so energizing and exciting to interact with the group at Frederick and all the people in the research community who have stepped up to help,” McCormick says.

Frank McCormick, Ph.D. Photo courtesy of UCSF.
The mission of the national lab, he says, is to facilitate and accelerate Ras drug discovery.
“The idea is for the national lab to derive structures of Ras proteins, and to develop reagents and cell lines and animal models and new screens for Ras inhibitors, and so on,” McCormick says. “These are things that may not be seen as innovative and wouldn’t be supported by grant funding to individual investigators, but they’re so essential to giving people the information they need to do the drug discovery.”
“Ras is one of the holy grails of cancer research and drug discovery,” Fesik says. “The idea is: ‘let’s put together our best ideas, our best technology; let’s take a risk and go after this very important target.’”
Fesik’s K-Ras Program is supported by the Vanderbilt-Ingram Specialized Program of Research Excellence (SPORE) in Gastrointestinal Cancer, the Lustgarten Foundation, the Alan Rittenberg Discovery Grant in Cancer Research and the NIH Director’s Pioneer Award. The Fesik drug discovery team is searching for compounds that block K-Ras and a number of other “undruggable” targets.
Fesik is a member of the Vanderbilt-Ingram Cancer Center, Orrin H. Ingram II Professor of Cancer Research, and professor of Biochemistry, Pharmacology, and Chemistry at Vanderbilt University.
